Professor
David Gatfield
Center for Integrative Genomics – UNIL Research in our group combines two main scientific themes, circadian rhythms and RNA biology.
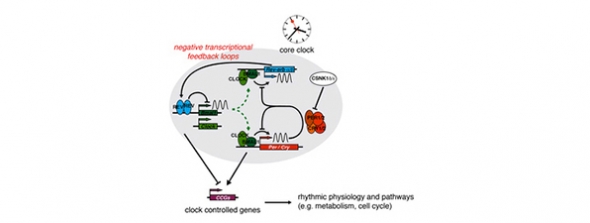
A considerable proportion of mammalian gene expression undergoes rhythmic oscillations driven by circadian clocks. While it is commonly thought that the majority of mRNA and protein rhythms is generated by cyclic transcription, there is accumulating evidence that post-transcriptional mechanisms make important contributions as well. Using mouse organs such as the liver and tissue culture cells as model systems, we investigate how mechanisms acting at the mRNA level participate in shaping the rhythmic transcriptome and proteome. In the past, a particular focus of our research has been on the activity of microRNAs in the context of circadian gene expression. Moreover, we have initiated additional research lines that aim to comprehensively investigate the regulation of translation around-the-clock (using ribosome profiling), and to discover RNA-binding proteins (RBPs) that function as regulators of core clock oscillator properties such as its period and phase. Using these RBPs as entry points, we wish to identify novel regulatory mechanisms at any level of RNA metabolism – from transcription to splicing, nuclear export, translation and degradation – that impact on rhythmic gene expression, metabolism and physiology.
Keywords
- Circadian clocks
- RNA biology
- translation
- microRNAs